Motivation
Reserving and projection methods are fitted on observed data, but
their practical value lies in how they would have performed at past
valuation dates. backtest() answers that question by hiding
the latest holdout calendar diagonals from a triangle,
refitting the model on the earlier portion, and comparing its projection
to the actuals that were withheld. This is calendar-diagonal hold-out
(rather than dev-period hold-out), because it simulates “what would the
model have said K months ago at the valuation date?”. The
cell-level metric follows the standard actuarial A/E convention,
,
where positive values flag under-projection (actual exceeded expected)
and negative values flag over-projection.
Basic usage
library(lossratio)
data(experience)
exp <- as_experience(experience)
tri_sur <- build_triangle(exp[cv_nm == "SUR"], cv_nm)
bt <- backtest(tri_sur, holdout = 6L)
print(bt)
#> <Backtest>
#> fit_fn : fit_lr
#> value_var : clr
#> holdout : 6 calendar diagonals
#> held-out : 123 cells
#> AEG : mean -13.05% / median -7.28%The returned object is a "Backtest" list with these key
slots:
-
aeg— per-celldata.table(cohort, dev, actual, pred, aeg, calendar_idx). -
col_summary— AEG aggregated bydev. -
diag_summary— AEG aggregated by calendar diagonal. -
masked— the triangle the fit was trained on (latest diagonals removed). -
fit— the fit object returned byfit_fn(anLRFitorCLFit).
summary(bt) prints the two summary tables alongside the
call metadata.
Validation coverage after masking
Masking the latest holdout diagonals shortens the
triangle’s lower-right edge. Chain ladder can only project as far as the
largest dev still observed in the masked data, so cells beyond that
range — the oldest cohorts at their latest dev — have no projection to
compare against. These unreachable cells are silently dropped, so
bt$aeg contains only cells where both an actual and a
finite projection exist.
Practical takeaway: as holdout grows, the validation set
shrinks fastest in the oldest cohorts’ late-dev region — exactly where
chain ladder relies on extrapolation (projection beyond the observed dev
range), so it is the area most in need of validation yet the first to
disappear.
Output interpretation
col_summary — systematic bias by development
period. A consistently signed AEG at a given dev signals a
structural mismatch between the model and that maturity. Early-dev
positive values usually reflect inflated link factors; late-dev values
flag tail miscalibration.
head(bt$col_summary, 8)
#> cv_nm dev n aeg_mean aeg_med aeg_wt
#> <char> <int> <int> <num> <num> <num>
#> 1: SUR 2 1 -0.2210212 -0.2210212 -0.2210212
#> 2: SUR 3 2 -0.6437701 -0.6437701 -0.6163919
#> 3: SUR 4 3 -0.3511641 -0.1162380 -0.3498190
#> 4: SUR 5 4 -0.3150824 -0.2157648 -0.3172642
#> 5: SUR 6 5 -0.4607816 -0.4015157 -0.4605004
#> 6: SUR 7 6 -0.3179501 -0.3459763 -0.3294385
#> 7: SUR 8 6 -0.3943149 -0.4362693 -0.3951618
#> 8: SUR 9 6 -0.3184528 -0.3718590 -0.3083244aeg_mean averages cell-level AEG, aeg_med
is the median, and aeg_wt = sum(actual - pred) / sum(pred)
is the exposure-weighted pooled A/E ratio minus 1. Comparing the three
columns flags whether a few large cells dominate (aeg_wt
very different from aeg_med) or the bias is uniform.
diag_summary — calendar-year effect. A
single bad diagonal in otherwise unbiased output points at a calendar
event (a rate change, claim handling shift, or one-off shock) that a
static fitter cannot see by construction.
bt$diag_summary
#> cv_nm calendar_idx n aeg_mean aeg_med aeg_wt
#> <char> <int> <int> <num> <num> <num>
#> 1: SUR 25 23 -0.1066364 -0.03677298 -0.07019853
#> 2: SUR 26 22 -0.1401734 -0.05189725 -0.11335460
#> 3: SUR 27 21 -0.1090998 -0.05853744 -0.10418242
#> 4: SUR 28 20 -0.1311208 -0.07720194 -0.12745825
#> 5: SUR 29 19 -0.1621096 -0.15960239 -0.16741775
#> 6: SUR 30 18 -0.1402920 -0.10632300 -0.16505109A monotone drift across calendar diagonals (as in the SUR example
above, where AEG becomes increasingly positive across
25, ..., 30) typically indicates that actuals on the latest
diagonals are running above what the earlier-cohort link factors imply,
i.e. a regime shift the static model has not absorbed.
aeg — cell-level outliers. For
diagnosing specific cohort × dev cells, inspect bt$aeg
directly:
head(bt$aeg, 5)
#> Key: <cv_nm>
#> cv_nm cohort dev value_actual value_pred aeg calendar_idx
#> <char> <Date> <int> <num> <num> <num> <int>
#> 1: SUR 2023-05-01 24 1.030446 1.156866 -0.109277544 25
#> 2: SUR 2023-06-01 23 1.175862 1.182519 -0.005629942 25
#> 3: SUR 2023-06-01 24 1.198728 1.292790 -0.072758288 26
#> 4: SUR 2023-07-01 22 1.105530 1.113031 -0.006738881 25
#> 5: SUR 2023-07-01 23 1.106120 1.118137 -0.010746990 26Plot demos
Four plot views are registered on "Backtest":
plot(bt, type = "col") # AEG by dev (point + dashed zero line)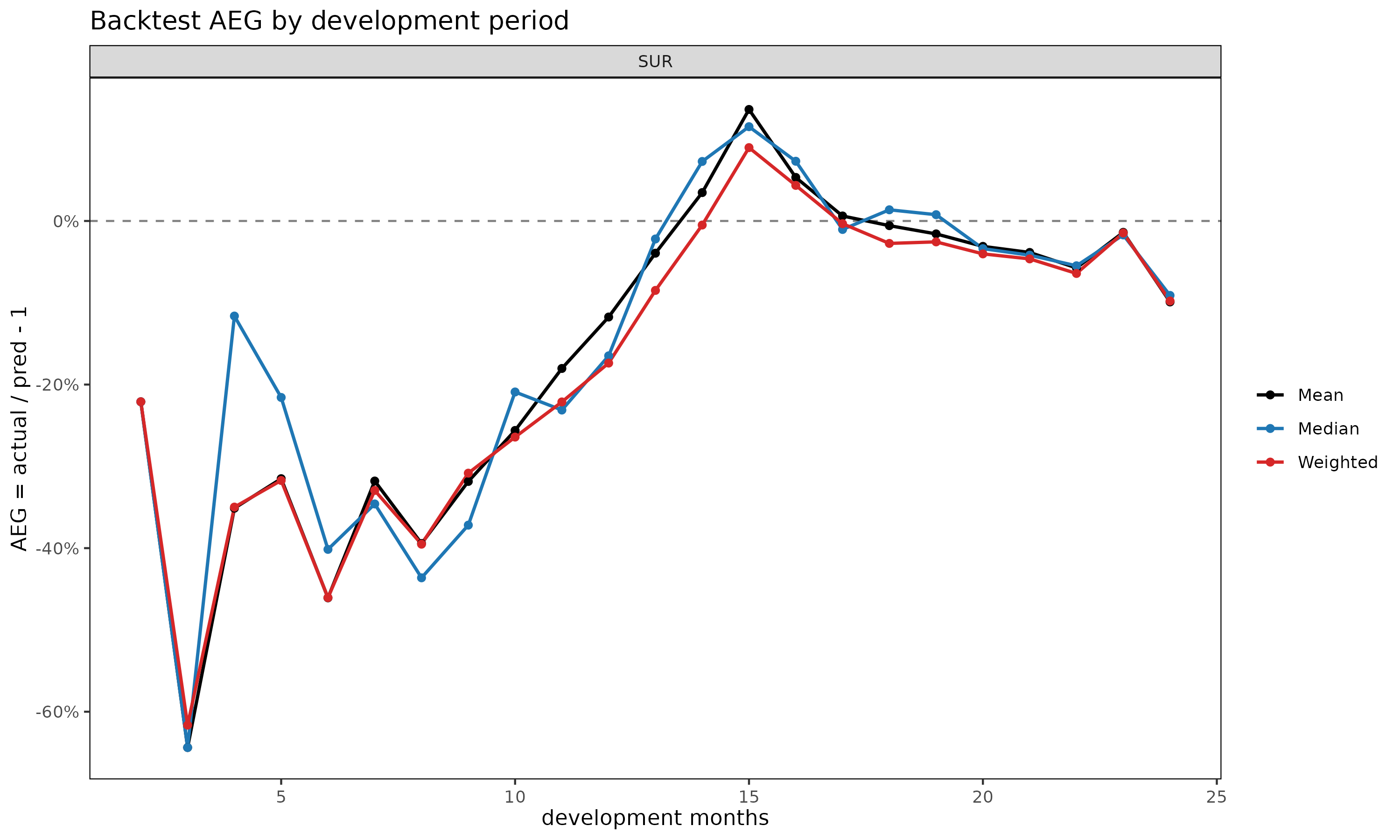
plot(bt, type = "diag") # AEG by calendar diagonal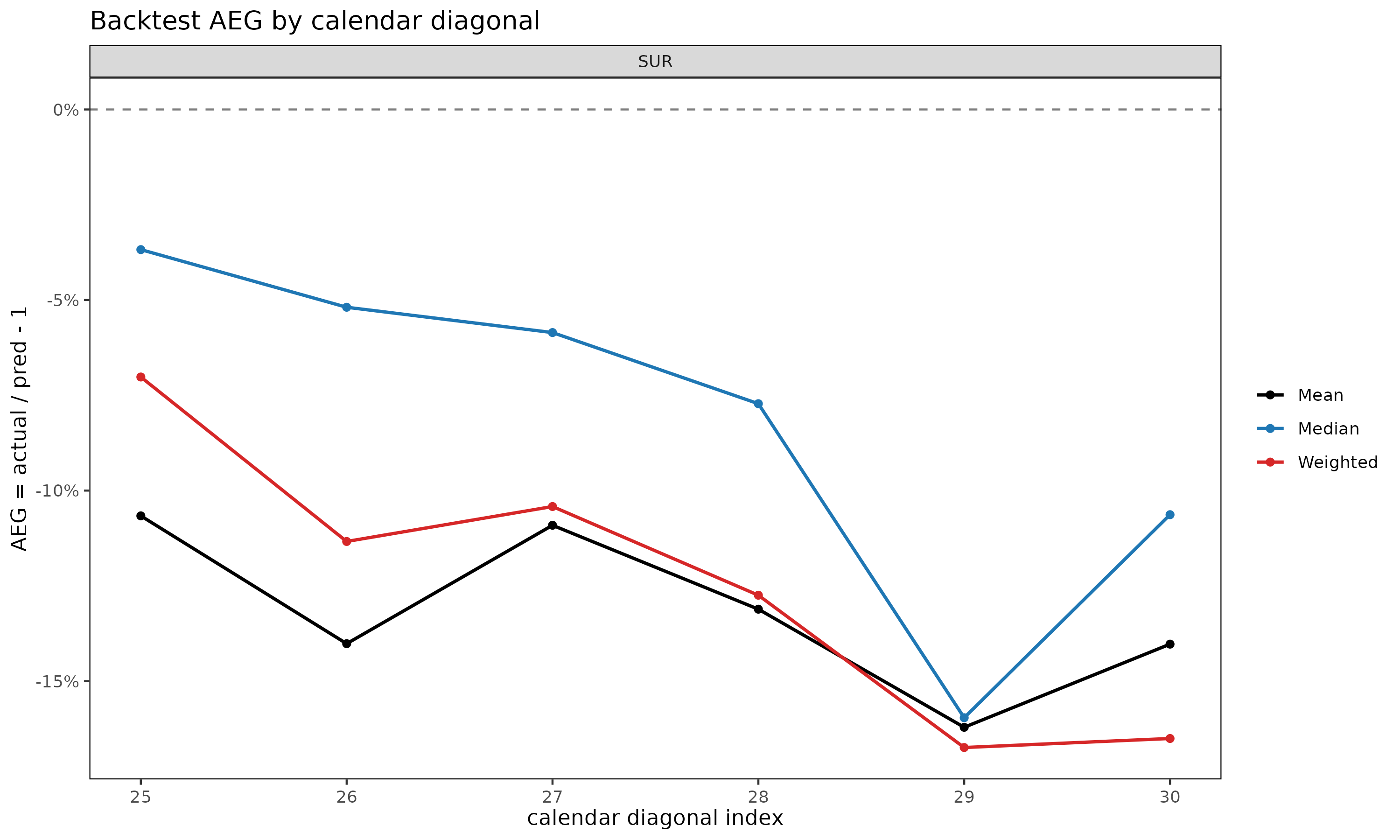
plot(bt, type = "cell") # per-cohort AEG trajectories over dev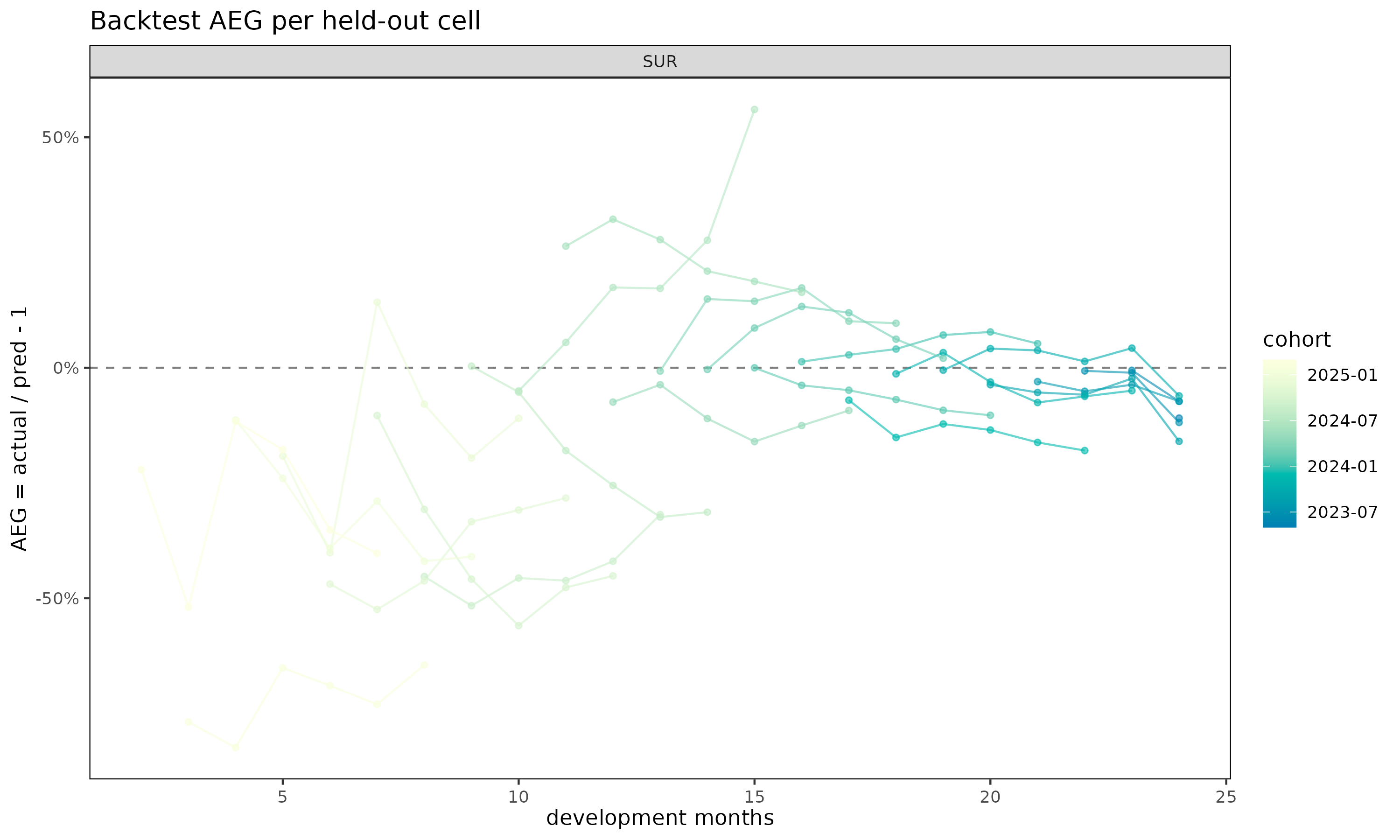
plot_triangle(bt) # diverging-color heatmap on the held-out wedge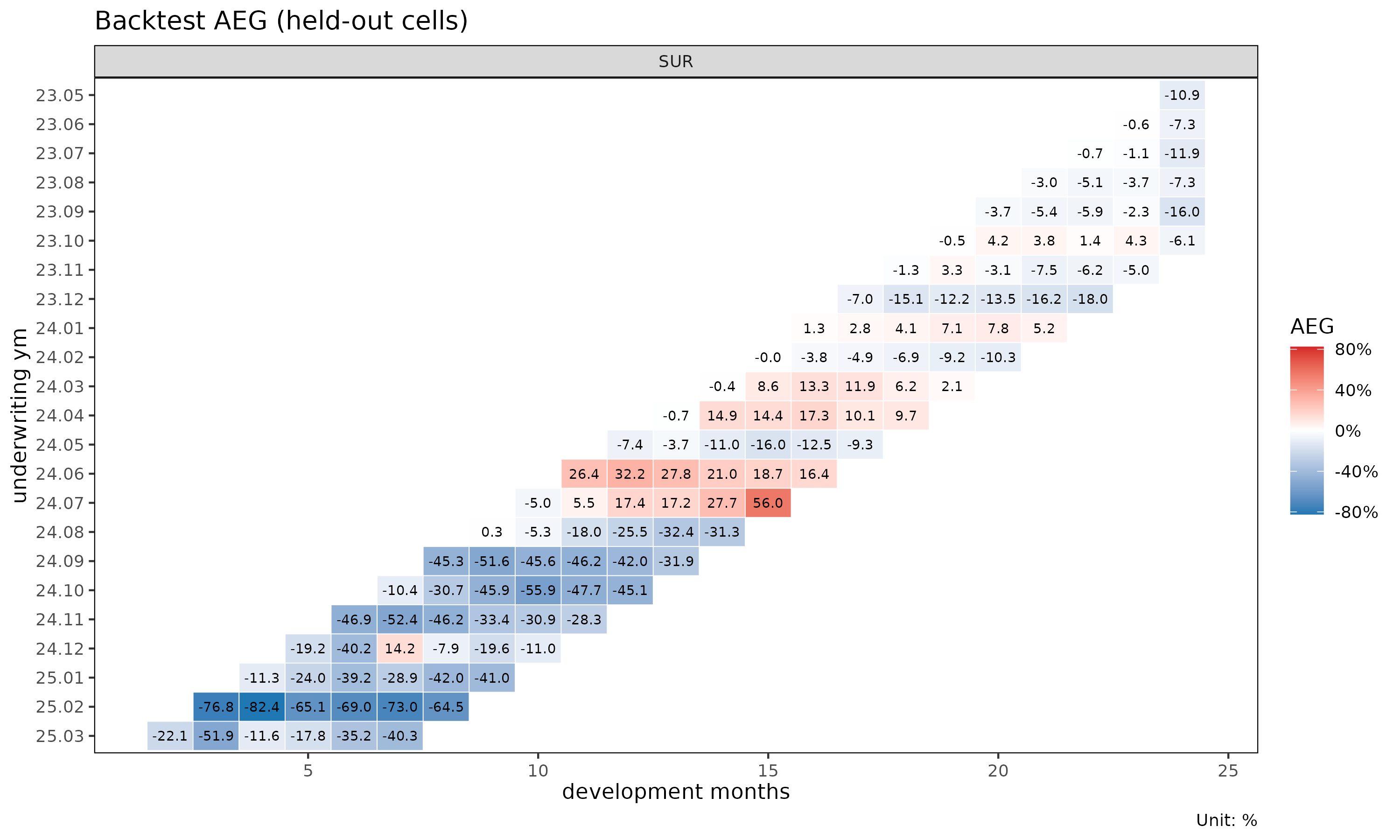
type = "col" is the right place to look for systematic
dev-period bias; type = "diag" reveals calendar-year drift;
type = "cell" exposes which cohorts contribute the bias;
plot_triangle() puts the cell-level AEG values on the same
triangular layout as plot_triangle() for the underlying
fit, with a red/blue diverging palette where red marks under-projection
(actual > pred).
Holdout selection
Choose holdout to balance two opposing effects:
- Too large: the masked triangle loses its latest experience, so the oldest cohorts have few or no reachable cells in their later dev periods. The validation set shrinks unevenly, biased toward early dev.
- Too small: the held-out wedge is just a thin diagonal band, which may not capture enough cells to reveal systematic patterns.
Typical choices are holdout = 6L (half-year) for monthly
triangles, or holdout = 12L (full year) for stronger
validation when the triangle has at least 24–30 diagonals of
history.
Choosing the fit function
The default fitter is fit_lr with
method = "sa" and value_var = "clr". The loss
ratio is unitless and dimension-free across cohorts of very different
volume, so aeg_mean and aeg_med carry a
consistent meaning across the triangle.
A note on
value_var.backtest(value_var = ...)is the score column — the column on which actual vs. predicted are compared cell-by-cell. It is not, in general, the same thing as thevalue_varargument to a chain-ladder fitter (which selects which column of the triangle to accumulate). Withfit_fn = fit_cl, the two coincide becausebacktest()forwardsvalue_varstraight through tofit_cl(). Withfit_fn = fit_lr, the fitter does not take avalue_varat all — it always projectscloss,crp, andclrjointly — andvalue_varhere only chooses which of those three projection columns onfit_lr$fullis compared against the held-out actuals:
value_var |
Compared column on fit_lr$full
|
|---|---|
"closs" |
loss_proj |
"crp" |
exposure_proj |
"clr" |
lr_proj |
The method argument selects the underlying loss-ratio
projection strategy: "sa" (stage-adaptive, the default)
blends exposure-driven projections before the maturity point with chain
ladder afterwards; "ed" is purely exposure-driven;
"cl" is the classical chain ladder applied to
clr.
bt_sa <- backtest(tri_sur, holdout = 6L, method = "sa") # default
bt_ed <- backtest(tri_sur, holdout = 6L, method = "ed")
bt_cl <- backtest(tri_sur, holdout = 6L, method = "cl")
bt_loss <- backtest(tri_sur, holdout = 6L, value_var = "closs")
bt_rp <- backtest(tri_sur, holdout = 6L, value_var = "crp")
print(bt_sa)
#> <Backtest>
#> fit_fn : fit_lr
#> value_var : clr
#> holdout : 6 calendar diagonals
#> held-out : 123 cells
#> AEG : mean -13.05% / median -7.28%Backtesting closs weights the result toward whichever
cohorts happen to be the largest at the held-out diagonals, which is
useful when monetary impact matters more than a normalized
comparison.
If you only need to project a single triangle column (e.g., raw
cumulative loss without forming a ratio), fit_fn = fit_cl
is also supported.
See also
-
vignette("chain-ladder")—fit_cl()reference. -
vignette("loss-ratio-methods")—fit_lr()and the"sa","ed","cl"methods. -
?backtest,?plot.Backtest,?plot_triangle.Backtest.
