fit_cl() is the dedicated chain ladder fit for a single
value column. Unlike fit_lr() (which projects loss /
exposure jointly to get loss ratio), fit_cl() projects one
cumulative metric forward and computes Mack-style standard errors per
cohort.
Basic usage
For brevity this vignette uses the SUR group only —
every step generalises to multi-group input.
library(lossratio)
data(experience)
exp <- as_experience(experience)
tri <- build_triangle(exp[cv_nm == "SUR"], group_var = cv_nm)
cl <- fit_cl(tri, value_var = "closs", method = "mack")
print(cl)
#> <CLFit>
#> method : mack
#> value_var : closs
#> weight_var : none
#> alpha : 1
#> sigma_method: min_last2
#> recent : all
#> use_maturity: FALSE
#> tail_factor : 1
#> groups : cv_nm
#> periods : 30value_var selects the cumulative column to project —
typically "closs" (cumulative loss) for reserving, or
"crp" (cumulative risk premium) for exposure
projection.
Method: basic vs Mack
Two estimation methods are available:
method |
What it computes |
|---|---|
"basic" |
Point projection only (selected age-to-age factors) |
"mack" |
Point projection + factor / process / parameter SE |
cl_basic <- fit_cl(tri, value_var = "closs", method = "basic")
cl_mack <- fit_cl(tri, value_var = "closs", method = "mack")
names(cl_basic)
#> [1] "call" "data" "method" "group_var"
#> [5] "cohort_var" "dev_var" "value_var" "full"
#> [9] "pred" "ata" "summary" "factor"
#> [13] "selected" "maturity" "alpha" "sigma_method"
#> [17] "weight_var" "recent" "use_maturity" "maturity_args"
#> [21] "tail" "tail_factor"
# Mack adds variance estimates to $full and $summary
head(cl_mack$summary)
#> cv_nm cohort latest ultimate reserve proc_se param_se
#> <char> <Date> <num> <num> <num> <num> <num>
#> 1: SUR 2023-04-01 2442597048 2442597048 0 0.0 0.0
#> 2: SUR 2023-05-01 2423543638 2600462324 176918686 270023.8 278555.7
#> 3: SUR 2023-06-01 3211045460 3634951626 423906166 461673.6 481436.4
#> 4: SUR 2023-07-01 2552396709 3106052713 553656004 217960839.6 130390124.6
#> 5: SUR 2023-08-01 2472997706 3159902325 686904619 235800277.9 139803599.9
#> 6: SUR 2023-09-01 2014222417 2712676349 698453932 230925350.1 124174149.3
#> se cv
#> <num> <num>
#> 1: 0.0 0.0000000000
#> 2: 387951.2 0.0001491855
#> 3: 667025.8 0.0001835034
#> 4: 253985259.7 0.0817710719
#> 5: 274129198.7 0.0867524279
#> 6: 262194082.1 0.0966551289method = "mack" enables the projection plot’s confidence
bands (show_interval = TRUE):
plot(cl_mack, type = "projection", show_interval = TRUE)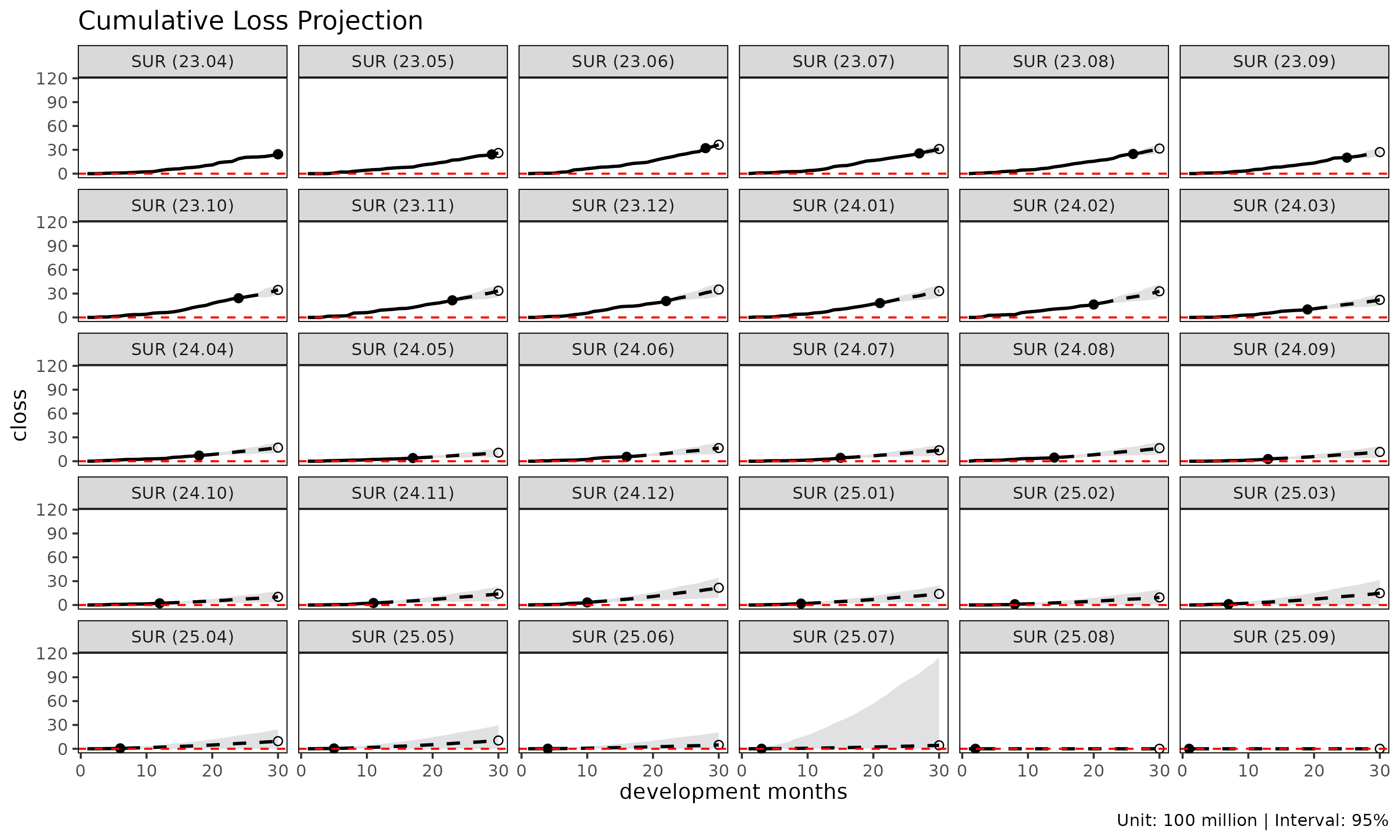
Tail factor
For triangles where the latest observed development period is still developing, an extrapolated tail factor estimates ultimate:
# Log-linear extrapolation from the selected ATA factors
cl_tail <- fit_cl(tri, value_var = "closs", method = "mack", tail = TRUE)
# Or supply a literal tail factor
cl_tail <- fit_cl(tri, value_var = "closs", method = "mack", tail = 1.025)The extrapolation fits
to projected factors and extends the projection by the cumulative
product of extrapolated
values. Disabled by default (tail = FALSE).
Maturity filtering
If selected ATA factors are volatile, restrict projection to the mature region only:
cl_mat <- fit_cl(
tri,
value_var = "closs",
method = "mack",
maturity_args = list(cv_threshold = 0.10, rse_threshold = 0.05)
)
cl_mat$maturity
#> Key: <cv_nm>
#> cv_nm ata_from ata_to ata_link mean median wt cv f f_se rse
#> <char> <num> <num> <char> <num> <num> <num> <num> <num> <num> <num>
#> 1: SUR 9 10 9-10 1.188 1.172 1.165 0.097 1.165 0.022 0.019
#> sigma n_obs n_valid n_inf n_nan valid_ratio
#> <num> <num> <num> <num> <num> <num>
#> 1: 1774.278 21 21 0 0 1maturity_args is forwarded to
find_ata_maturity().
Variance components (Mack)
fit_cl(method = "mack") decomposes the projection
variance into:
-
proc_se— process variance, from (residual link variance per development period). -
param_se— parameter variance, from the uncertainty of the selected age-to-age factors . -
se— total standard error, . -
cv— coefficient of variation,se / value_proj.
summary(cl_mack)
#> cv_nm cohort latest ultimate reserve proc_se param_se
#> <char> <Date> <num> <num> <num> <num> <num>
#> 1: SUR 2023-04-01 2442597048 2442597048 0 0.0 0.0
#> 2: SUR 2023-05-01 2423543638 2600462324 176918686 270023.8 278555.7
#> 3: SUR 2023-06-01 3211045460 3634951626 423906166 461673.6 481436.4
#> 4: SUR 2023-07-01 2552396709 3106052713 553656004 217960839.6 130390124.6
#> 5: SUR 2023-08-01 2472997706 3159902325 686904619 235800277.9 139803599.9
#> 6: SUR 2023-09-01 2014222417 2712676349 698453932 230925350.1 124174149.3
#> 7: SUR 2023-10-01 2422172254 3464336723 1042164469 276909538.1 163800531.1
#> 8: SUR 2023-11-01 2157147612 3350616805 1193469193 347646646.1 180286627.2
#> 9: SUR 2023-12-01 2062030017 3510350121 1448320104 379204803.4 195291838.6
#> 10: SUR 2024-01-01 1803809914 3316423447 1512613533 371903391.7 185291186.8
#> 11: SUR 2024-02-01 1627213157 3293904272 1666691115 406210629.2 191768888.6
#> 12: SUR 2024-03-01 1006624213 2212909862 1206285649 348109439.7 131371151.5
#> 13: SUR 2024-04-01 707083237 1712964993 1005881756 316686848.8 103164315.5
#> 14: SUR 2024-05-01 398857325 1069653556 670796231 262671178.8 65778222.8
#> 15: SUR 2024-06-01 558855276 1654603718 1095748442 342800640.3 103939643.9
#> 16: SUR 2024-07-01 423131371 1378042306 954910935 336548946.6 89486750.7
#> 17: SUR 2024-08-01 457705980 1642689597 1184983617 387322897.0 109347246.8
#> 18: SUR 2024-09-01 278007651 1166380310 888372659 360265104.1 81491440.0
#> 19: SUR 2024-10-01 214811381 1027414214 812602833 358796880.9 74015042.1
#> 20: SUR 2024-11-01 251273971 1400108561 1148834590 451728990.6 105050619.0
#> 21: SUR 2024-12-01 322678179 2168358632 1845680453 619598522.3 171876903.1
#> 22: SUR 2025-01-01 179253475 1403314539 1224061064 523388251.8 114399770.9
#> 23: SUR 2025-02-01 100816665 954214626 853397961 497168458.4 84734261.7
#> 24: SUR 2025-03-01 111279087 1488227953 1376948866 843027195.9 163376024.6
#> 25: SUR 2025-04-01 55914454 958667529 902753075 751897091.4 113958278.9
#> 26: SUR 2025-05-01 41578391 1041506115 999927724 978637599.4 147793339.1
#> 27: SUR 2025-06-01 14997314 484991215 469993901 813441277.3 81273429.5
#> 28: SUR 2025-07-01 6232031 436725855 430493824 5630776358.8 495928426.0
#> 29: SUR 2025-08-01 0 0 0 0.0 0.0
#> 30: SUR 2025-09-01 0 0 0 0.0 0.0
#> cv_nm cohort latest ultimate reserve proc_se param_se
#> <char> <Date> <num> <num> <num> <num> <num>
#> se cv
#> <num> <num>
#> 1: 0.0 0.000000e+00
#> 2: 387951.2 1.491855e-04
#> 3: 667025.8 1.835034e-04
#> 4: 253985259.7 8.177107e-02
#> 5: 274129198.7 8.675243e-02
#> 6: 262194082.1 9.665513e-02
#> 7: 321728932.9 9.286884e-02
#> 8: 391613915.1 1.168782e-01
#> 9: 426538609.2 1.215089e-01
#> 10: 415505663.8 1.252873e-01
#> 11: 449201938.9 1.363737e-01
#> 12: 372073328.0 1.681376e-01
#> 13: 333066714.3 1.944387e-01
#> 14: 270782057.7 2.531493e-01
#> 15: 358211848.8 2.164940e-01
#> 16: 348242834.9 2.527084e-01
#> 17: 402462230.4 2.450020e-01
#> 18: 369366755.4 3.166778e-01
#> 19: 366351509.1 3.565763e-01
#> 20: 463783045.7 3.312479e-01
#> 21: 642996110.9 2.965359e-01
#> 22: 535744873.7 3.817711e-01
#> 23: 504337556.8 5.285368e-01
#> 24: 858712162.8 5.770031e-01
#> 25: 760483875.8 7.932718e-01
#> 26: 989734521.0 9.502916e-01
#> 27: 817491334.5 1.685580e+00
#> 28: 5652573520.7 1.294307e+01
#> 29: 0.0 NA
#> 30: 0.0 NA
#> se cv
#> <num> <num>Reserve plot
type = "reserve" shows reserve per cohort with optional
error bars (Mack only):
plot(cl_mack, type = "reserve", conf_level = 0.95)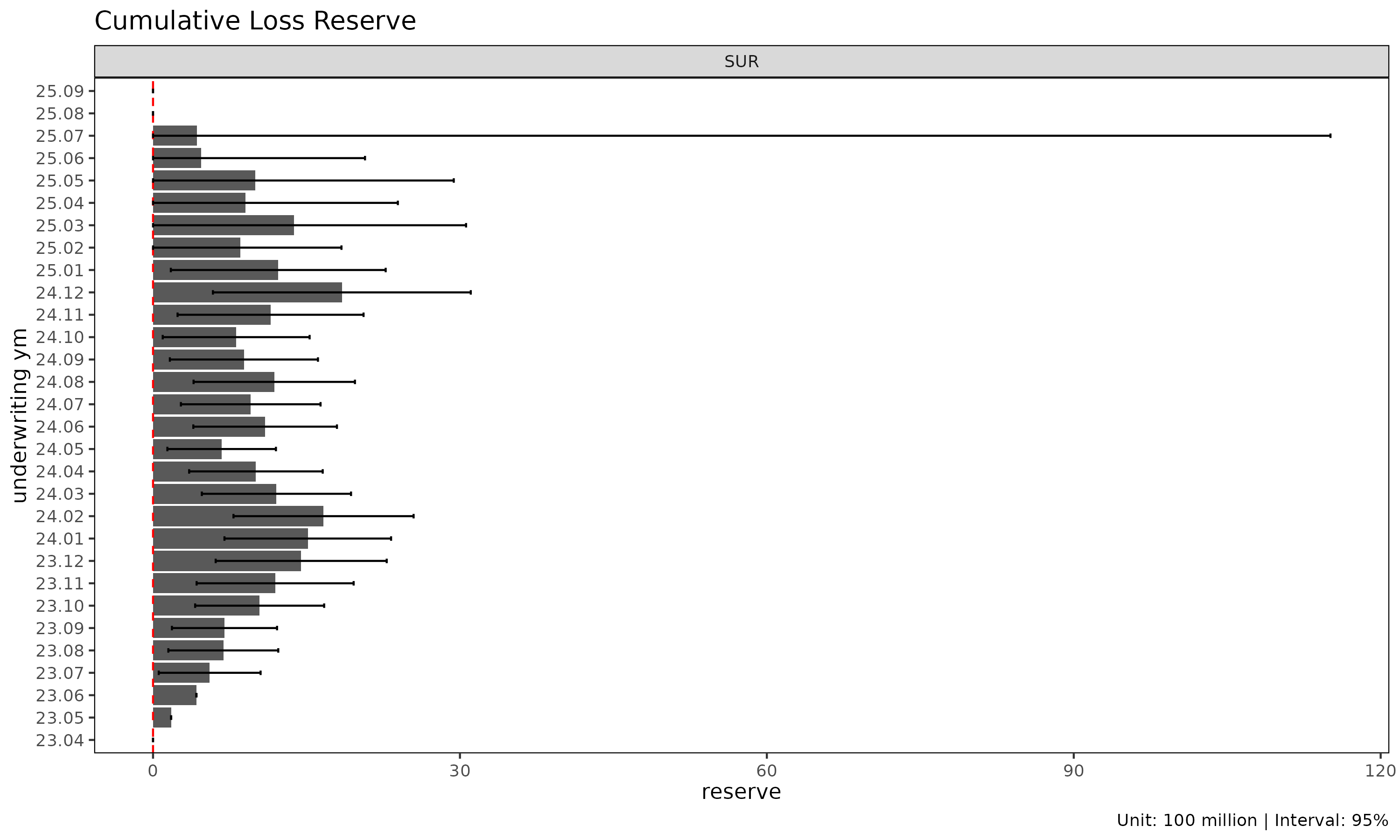
Triangle visualisation
plot_triangle() displays the cohort × dev cells as a
heatmap, distinguishing observed cells from projected:
plot_triangle(cl_mack, what = "full") # observed + projected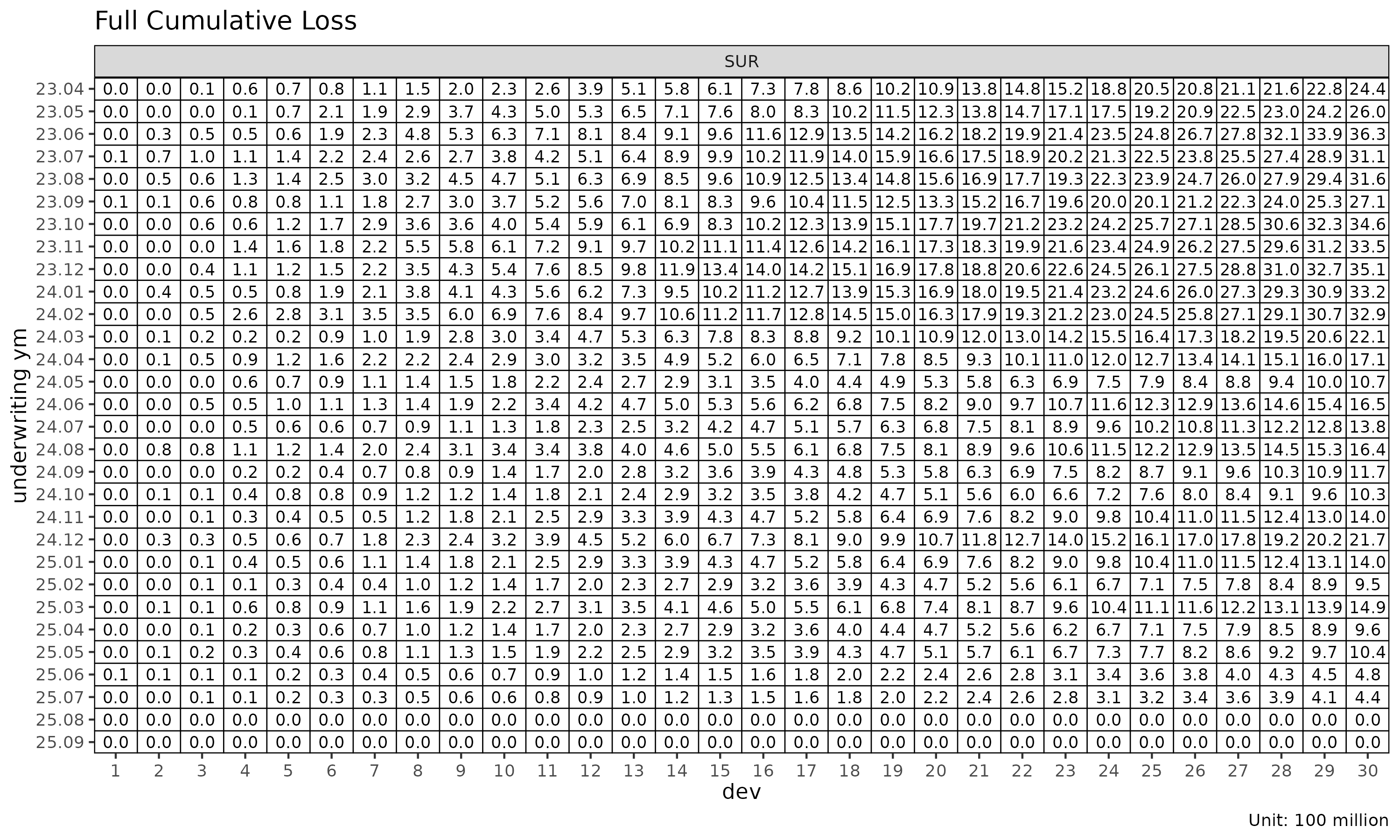
plot_triangle(cl_mack, what = "pred") # projected only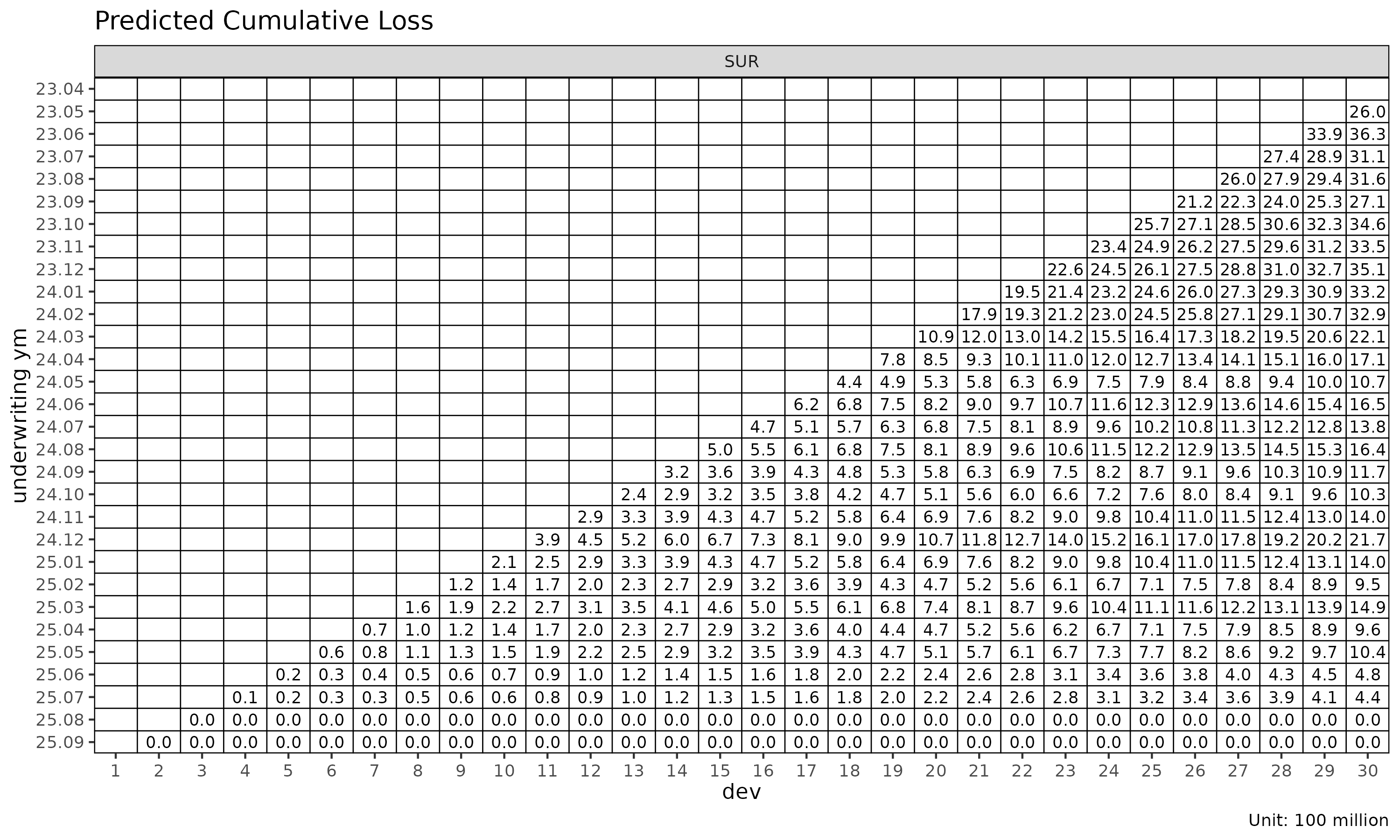
plot_triangle(cl_mack, what = "data") # observed only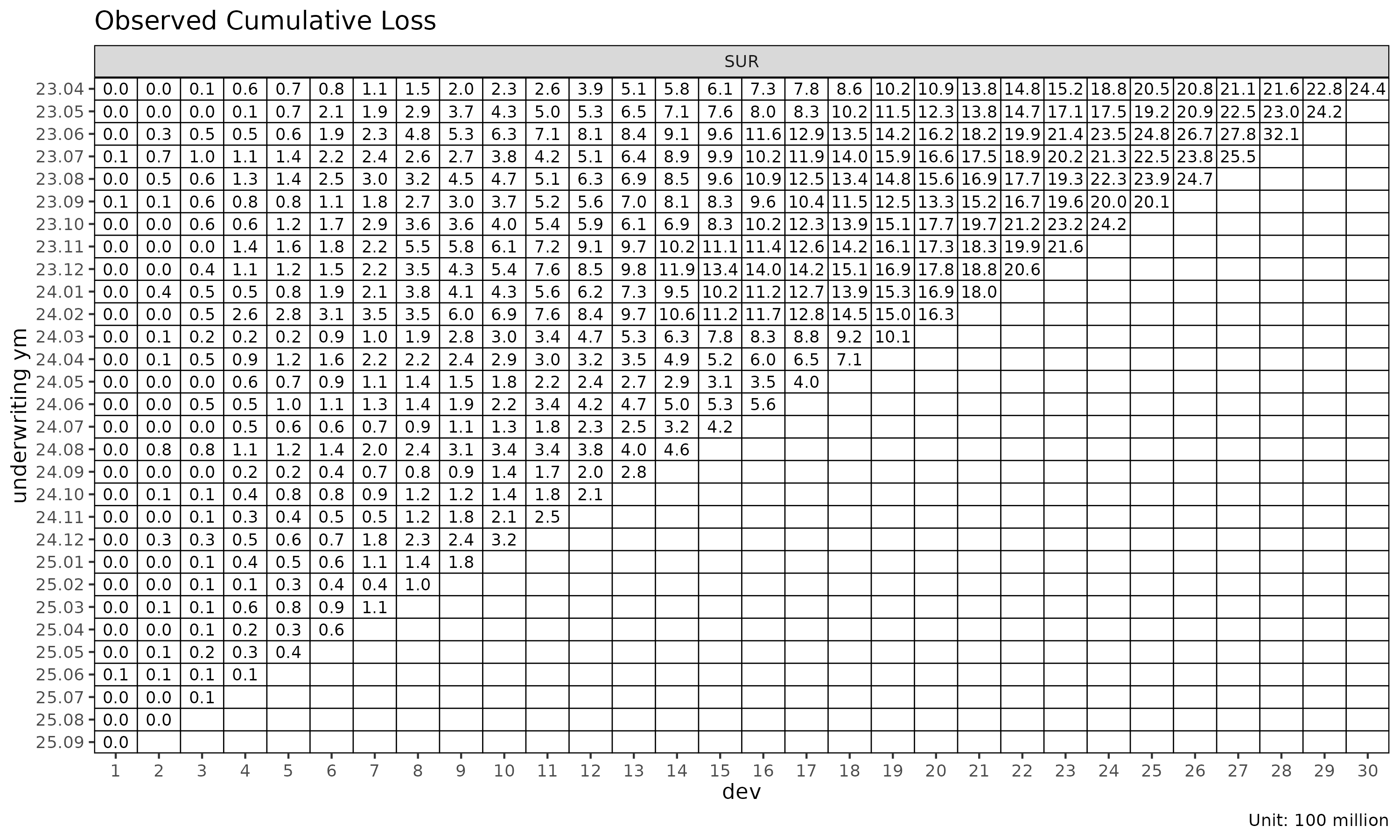
The label_style = "cv" mode shows coefficient of
variation per cell, useful for spotting unreliable cells:
plot_triangle(cl_mack, label_style = "cv")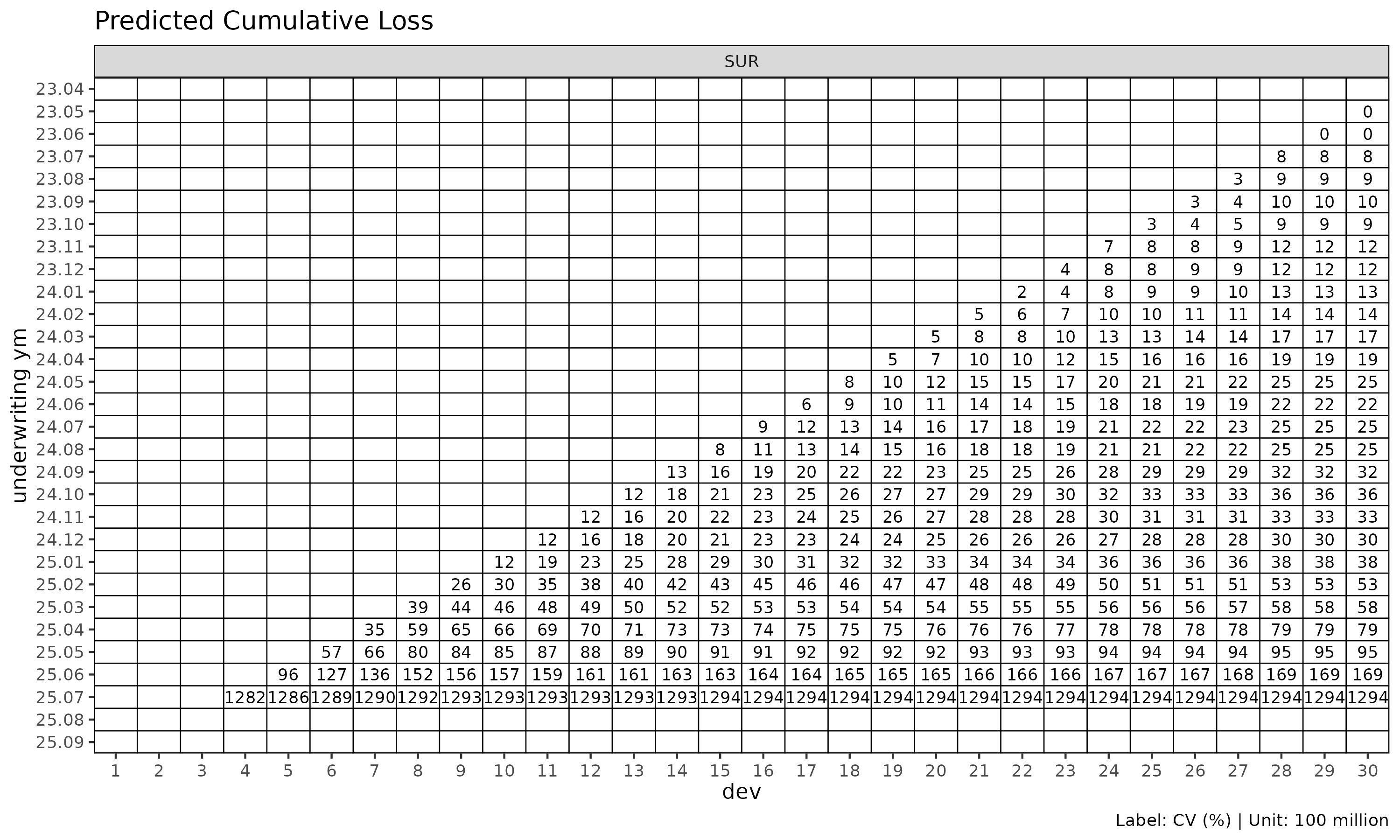
plot_triangle(cl_mack, label_style = "se")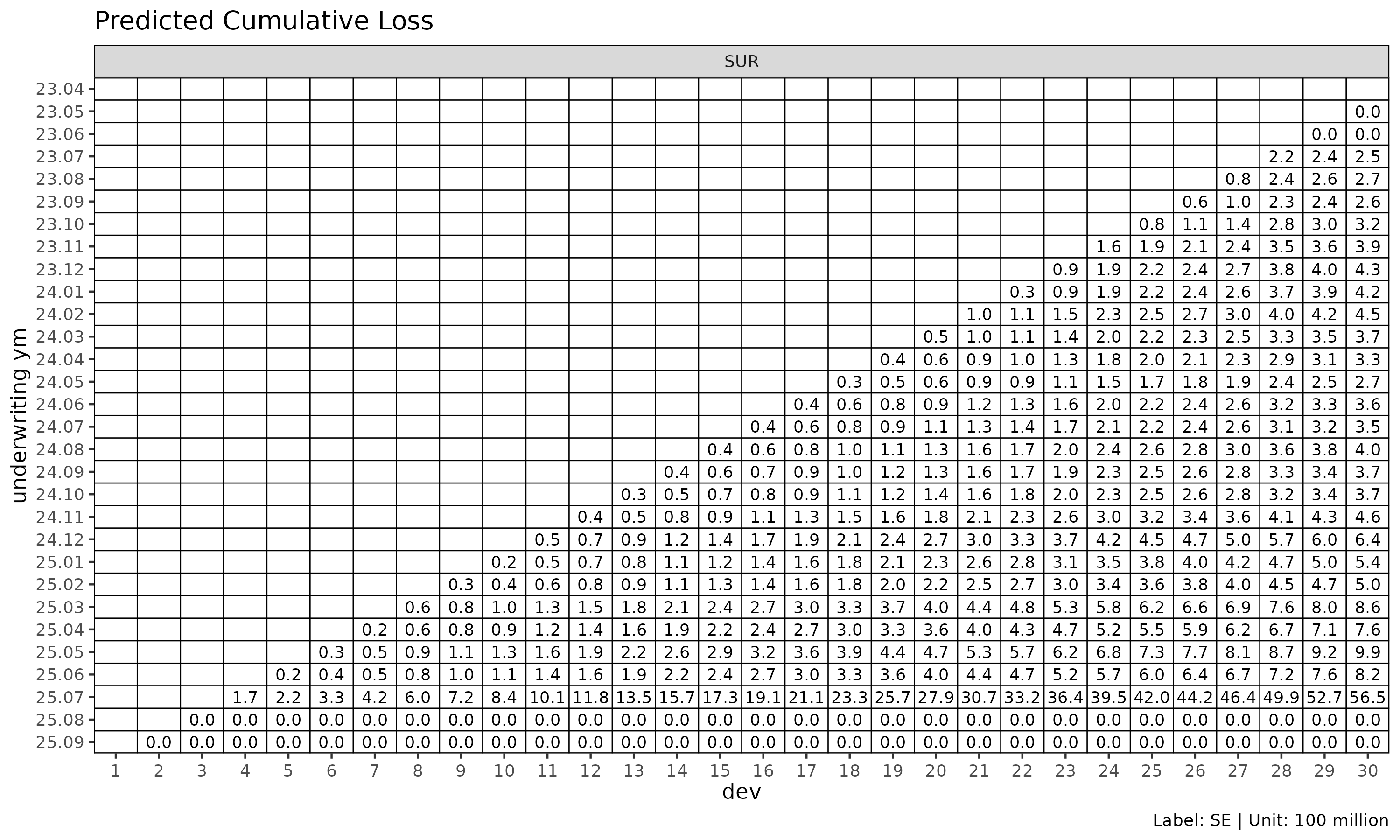
plot_triangle(cl_mack, label_style = "ci")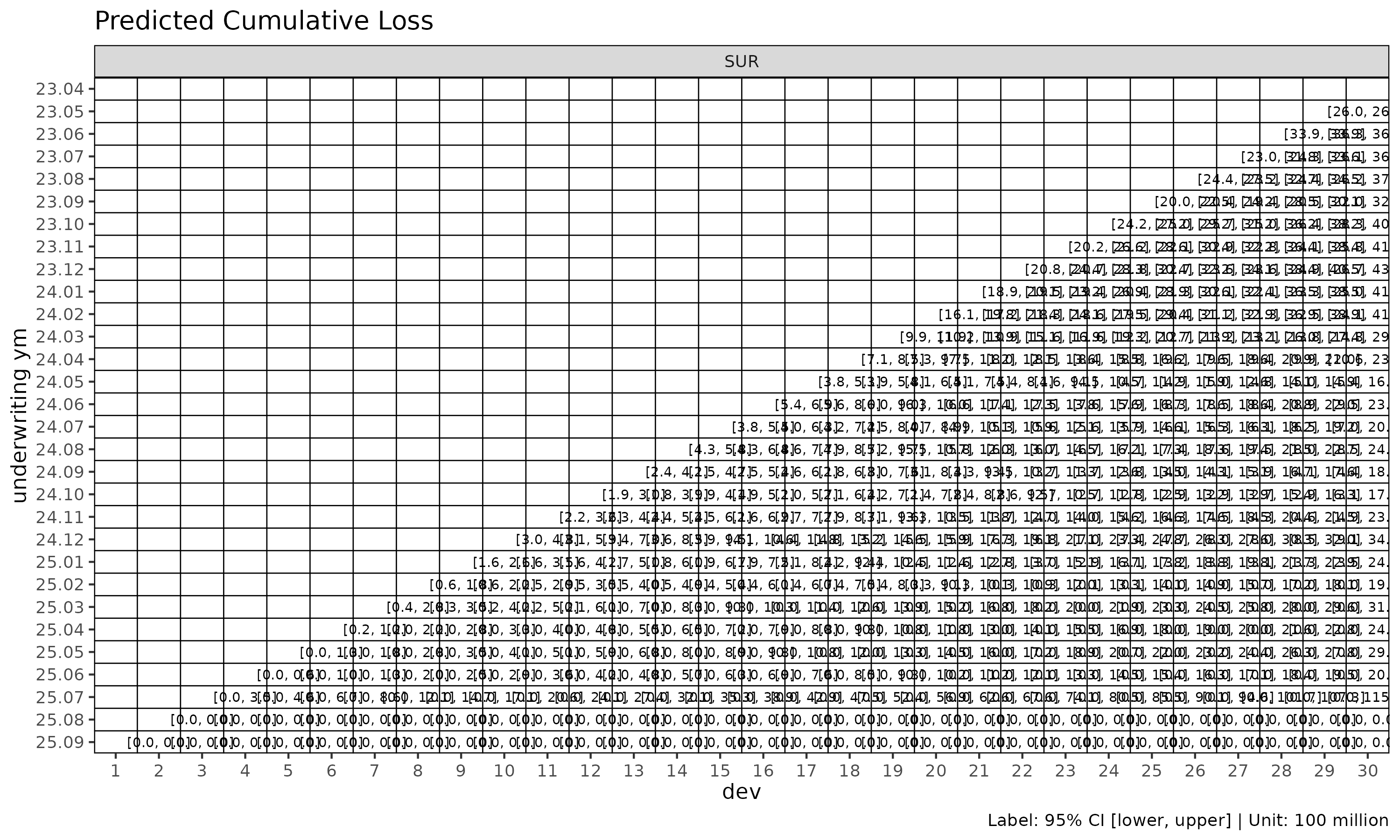
Sigma extrapolation methods
Mack variance requires
at all development links, including the last where it cannot be
estimated directly. sigma_method controls the
extrapolation:
sigma_method |
Behaviour |
|---|---|
"min_last2" |
(default) min of the last two estimable values — conservative |
"locf" |
Last observation carried forward |
"loglinear" |
Log-linear extrapolation from the observed sequence |
fit_cl(tri, value_var = "closs", method = "mack", sigma_method = "loglinear")
#> <CLFit>
#> method : mack
#> value_var : closs
#> weight_var : none
#> alpha : 1
#> sigma_method: loglinear
#> recent : all
#> use_maturity: FALSE
#> tail_factor : 1
#> groups : cv_nm
#> periods : 30See also
-
vignette("loss-ratio-methods")— when to usefit_lr()instead. -
vignette("triangle-diagnostics")—summary(),find_ata_maturity(), ata diagnostic plots. -
?fit_cl,?find_ata_maturity,?fit_ata.
